Story by Pippa Wysong. Source: Canadian Light Source

Biochemists from McGill University are getting a good look at just how a specific enzyme that is part of the human immune system interacts with a certain group of bacteria that are described as gram-negative.
Researchers around the world “have been studying the enzyme, known as AOAH, for more than 30 years. This is the first time anyone has been able to see exactly what it looks like,” according to Bhushan Nagar, an associate professor of biochemistry at McGill University in Montreal.
More than that, the 3D images captured a moment in time which shows just how AOAH inactivates a toxic molecule that is commonly part of various gram-negative bacteria. The research was conducted at the Canadian Light Source.
Numerous types of gram-negative bacteria exist throughout the environment. While some are harmless, many cause a variety of human illnesses, says Nagar. For example, several species such as E. coli and Salmonella, cause food borne illness. Others cause infections such as pneumonia, meningitis, bloodstream infections or gonorrhea.
Some types of gram-negative bacterial infections are demonstrating drug resistance and are increasingly difficult to treat, making understanding how the human immune system interacts with them even more important.
When an infection develops, it generally triggers inflammation through a toxic component of the outer shell of gram-negative bacteria, called lipopolysaccharide (LPS) – also commonly referred to as an endotoxin.
“Inflammation is a protective response by the immune system when fighting infections,” says Nagar.
It’s what causes fever, aches, swelling and sometimes redness on the skin. But there is a fine balance. Too much inflammation can cause a person to become sicker and even die, while too little won’t help kill off the bacteria causing the infection. The immune system controls this problem by dampening inflammation after the initial acute response. That way, the inflammation doesn’t cause excessive damage.
After the infection is cleared, the system needs to reset itself so it will be prepared to fight new infections. This job falls into the hands of the AOAH enzyme. With the gram-negative bacterium, it was found that the AOAH targets, binds to, and removes two specific regions on LPS, rendering it ineffective and causing the system to reset.
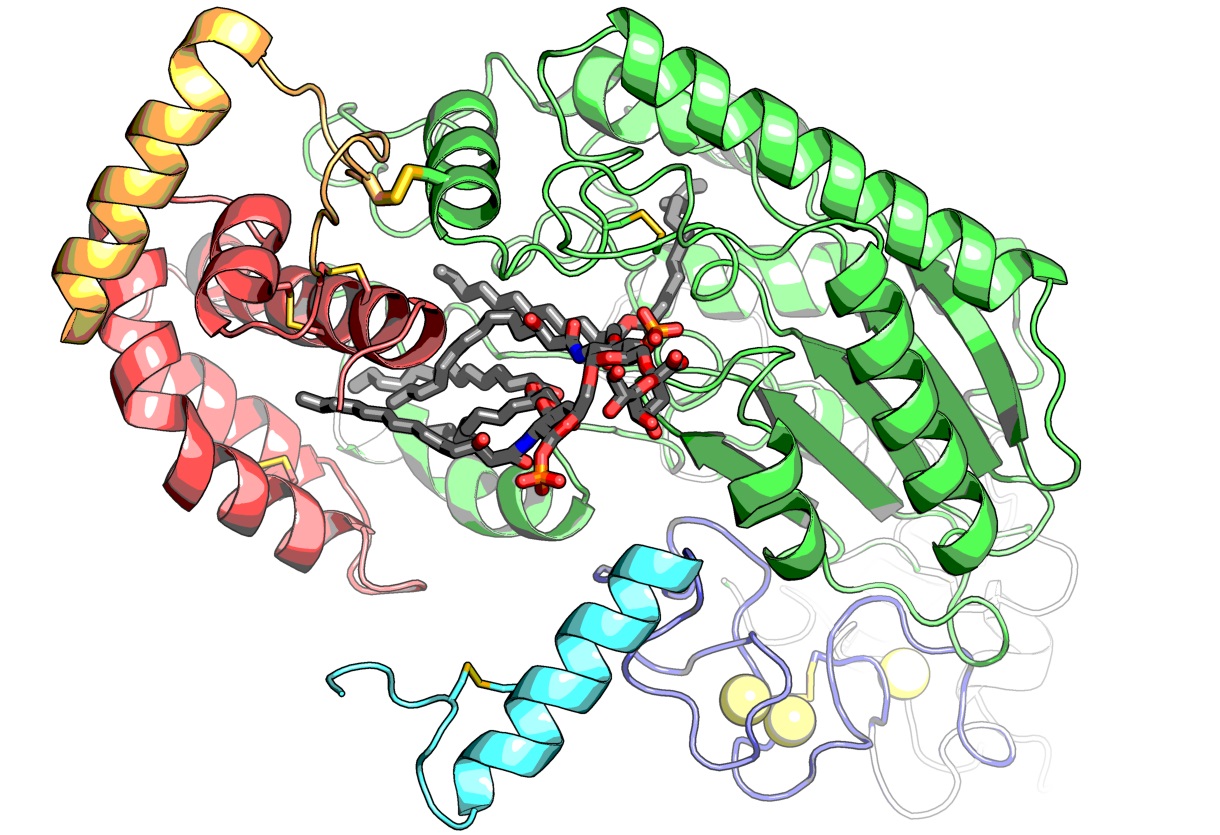
The researchers were able to see the images by using a crystallized sample of AOAH bound to a portion of the LPS. The results allowed for a detailed look at the molecular structure of AOAH and clearly showed which parts of its surface interacted with the toxin.
“We could actually see the molecular mechanism the enzyme uses to clip off parts of LPS to inactivate it,” says Nagar.
The findings were published in the Proceedings of the National Academy of Sciences earlier this year.
The images provide a map to the AOAH enzyme and opens the door to other directions of research. Some forms of AOAH are associated with certain diseases such as asthma and chronic sinusitis. Knowing its full structure will help researchers better understand what its disease-causing variations are like and could provide details needed to develop new treatments for infections or approaches to prevent them.
Alexei Gorelik, Katalin Illes, and Bhushan Nagar. “Crystal structure of the mammalian lipopolysaccharide detoxifier.” Proceedings of the National Academy of Sciences (2018): 201719834. DOI: 10.1073/pnas.1719834115
April 10, 2018
